Precision Medicine Solution Accelerates GATK Pipeline by 36x while Reducing Costs by 10x and Delivering 99.9% Accuracy over CPU-Only Solutions

Next-Gen Genome Sequencing (NGS) utilizes HPE GPU servers, NVIDIA Parabricks software, Weka parallel file system and Groupware AI Labs
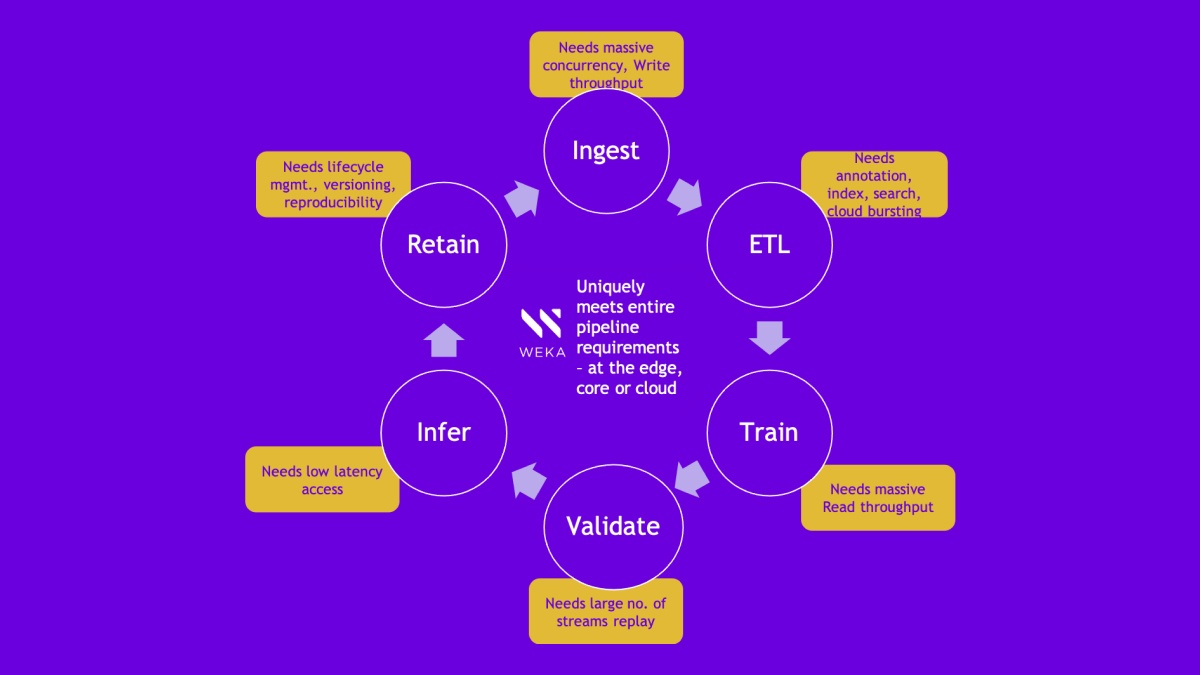
The COVID-19 pandemic has put the spotlight on whole genomes, whole exomes, germline DNA, somatic data sets, and RNA sequencing experiments using Next-Generation Sequencing (NGS) technology that is at the forefront of our attempts to understand, fight, and eliminate pathogens. When it comes to COVID-19 and genome samples from patients, NGS has discovered eight strains and 30,000 base pairs, confirming that the mutations are not much different from the original strain. NGS is an essential step toward gaining a better understanding of the root cause diseases, identifying new biomarkers associated with specific diseases, understanding new drug candidates, and personalizing treatments based on patient’s genetics. A whole human genome at 30X coverage can require several hundred gigabytes of storage during the alignment and variant calling stages, and it can take more than 30 hours to process this data using CPU-only solutions. As a result, there is a computational bottleneck when processing thousands of genomes or when a patient in critical condition is waiting in a clinical setting. HPE, NVIDIA, WekaIO (Weka), and Groupware have partnered to launch an accelerated NGS solution that overcomes these challenges by using GPU computing with NVIDIA Parabricks and the NVMe flash-based WekaFS parallel filesystem to accelerate the Genome Analysis Tool Kit (GATK).
Genomics Industry in Transition – the Focus is Moving from Sequencing to Interpretation
A key trend with genomics is that the focus has shifted from primarily sequencing new genotypes to interpretation of the data sets. In terms of technology, this means a significant shift to machine learning and GPU-based workloads, developing AI models, and leveraging large amounts of historical data sets to find new insights. NGS has progressed to where the cost of whole-genome processing has dropped below $1,000. This has resulted in a significant increase in genome processing as well as exponential data growth.
NGS Workflows
In a genomics workflow, the first step is data generation from sequencing instruments, such as the ones from Illumina. This data is sent to storage devices, which are typically Windows-based servers. These servers generally push the data to high-performance computing (HPC) cluster storage, which is where WekaFS integrates. WekaFS can connect directly to the instrumentation for data ingest, provide data to the HPC processing cluster, and store it long-term as part of a massive data store for collaboration.
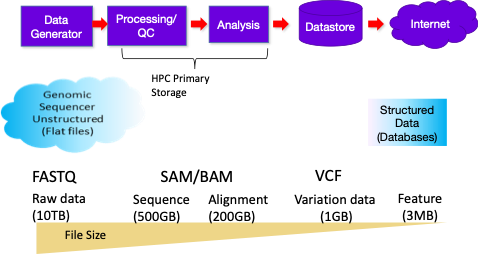
Genomics data is massive in its raw form because of image files generated by the sequencing instruments. Inside the sequencer, the images are converted to a FASTQ file (called BCL2FASTQ). Based on the desired accuracy, these sequencing files can be up to 10 terabytes. As the data is processed, it is converted into sequence alignment map (SAM) or Binary alignment map (BAM) files. These files are much smaller in size, and in fact, BAM files are compressed. Researchers take the resulting information, compare it to a reference genome, and record the differences in text files with variant call format (VCF).
Business Challenges with NGS
Here are the key business challenges that NGS presents:
- Life sciences organizations are overwhelmed by the rate of data growth, which is:
- doubling every 7 months – amounting to a total of40 exabytes by 2025
- straining research budgets
- There is a highly competitive race to get products to market
- performance matters
- Managing and protecting data at this scale is a nightmare
- data needs to be secured and protected
- there is too much data, more than can be backed up
The Solution Built on Weka AI
Fortunately, there is a solution that improves performance by 36x and reduces cost by 10x when compared to traditional CPU-based solutions and also delivers 99.9% accuracy.
This solution leverages the Weka AI blueprint to architect, deploy, test, and operationalize an end-to-end solution. The solution stack consists of:
- HPE Apollo 6500 GPU-accelerated servers
- 4 Apollo servers, each with 8 NVIDIA Tesla V100 GPUs, were used to create a compute cluster for distributed NGS.
- NVIDIA Parabricks software
- The NVIDIA Parabricks germline pipeline software accelerates the original Broad Institute GATK Best Practices pipeline using NVIDIA GPUs.
- Weka
- A 20 TB high-performance WekaFS cluster is used for these experiments on 8 HPE Proliant DL360 storage servers with NVMe for a flash tier.
- WekaFS provides hybrid workflows where AWS cloud can be leveraged for Cloud bursting and Disaster recovery.
- Groupware AI Labs
- Groupware expedites the NGS customer AI journey. Whether you are currently using AI or are just starting to formulate your AI policy, AI Labs provides comprehensive help from identifying business objectives all the way to racking, stacking, and operationalizing your AI infrastructure.
Key Performance Indicators (KPIs) for the Solution
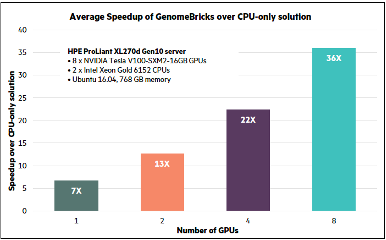
The combined solution leverages GPU-accelerated NVIDIA Parabricks GATK pipeline software running on HPE GPU compute servers. The WekaFS cluster ensures maximum throughput and the lowest latency when feeding data for this distributed pipeline to 4 Apollo servers.
- For 43x coverage level of Human NA12878 whole genome sequence, which provided a 76GB dataset, 464 GB of metadata, and 1 GB of output
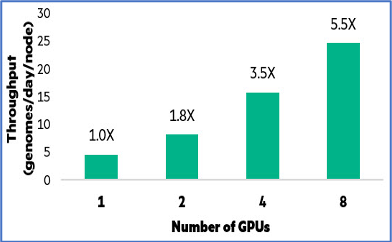
- Single job performance on a single node was able to linearly scale by 5.5x from 1 GPU to 8 GPUs
- When multiple concurrent jobs were run on a single node, they performed 6x better; and on 4 nodes, over 24x better compared to single job performance on single GPU.
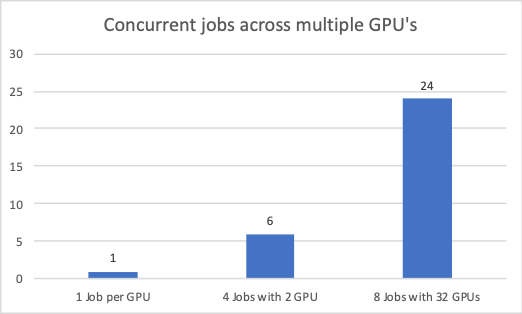
- Reduced number of CPU servers, combined with savings in storage, networking, power, space, and cooling can result in 9x-10x TCO reduction.
Conclusion
To summarize, this precision medicine solution overcomes the challenges posed by NGS workflows, and enables customers to quickly process genomics data, to be the first to insights and the first to market.
For more information, please see:
- HPE: Accelerated throughput of Next-Gen Sequencing
- HPE Tech Brief: HPE Apollo 6500 System with WekaIO File System Advances Next-Generation Sequencing
- Groupware Technology Solutions Brief Accelerated AI Genomic Discoveries with Parabricks and WekaIO Powered by HPE and NVIDIA
- WekaFS™ for Genomics: https://www.weka.io/resources/solution-briefs/accelerate-genomic-discovery-petagene-western-digital-and-wekaio
Authored by: Shailesh Manjrekar
Contributors:
Haruna Cofer, HPE
Chris Davidson, HPE
Joe Handzik, NVIDIA
Robert Clark, NVIDIA
Amrinderpal Singh Oberai, Groupware